Development of Potential COVID-19 Vaccine and Serological Assay
ABOUT THIS STUDY
Our group has explored the use of genomic RNA/phage display libraries derived from primary human malignant melanoma cells as a means of identifying antibody detectable targets on cancer cells (cancer vaccines or antibody guided therapeutics). In this approach, we isolate and affinity-column immobilize the IgG fraction from patient serum before and after immune therapy for melanoma, and expose the immobilized antibodies to bacteriophage expressing approximately 2x109 overlapping cDNA sequences of paired (same patient derived plasma and cancer cells) melanoma genomic RNA. Phage, expressing melanoma cDNA express the proteins/peptides on their capsid are “recognized” by the immobilized antibodies are retained in the column, and subsequently eluted for DNA sequencing. Comparison of the DNA profiles of the eluted phage using pre-immunotherapy and post-immunotherapy patient sera will reveal emergence of new antibodies (post-immunotherapy gain of antibodies) against proteins of potential interest for melanoma targeting. In the current proposal, we hypothesize that reacting COVID serum from patients that have recovered from COVID infection and compare to non-infected self-serum (if available) and control healthy volunteer serum (available in our lab) may identify protein targets that have developed as a result of the COVID infection and could be useful in the development of a COVID vaccine as well as a serologic test for anti-COVID immunity.
Participation eligibility
Participant eligibility includes age, gender, type and stage of disease, and previous treatments or health concerns. Guidelines differ from study to study, and identify who can or cannot participate. There is no guarantee that every individual who qualifies and wants to participate in a trial will be enrolled. Please contact the study team to discuss whether or not you are eligible to participate in a study.
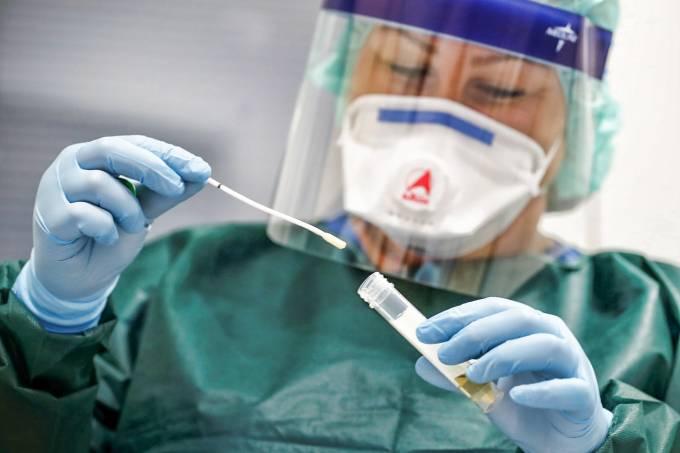
Readmore: COVID-19 Test Kit